Orienting Single Molecules
Although physicists have developed many methods for observing the average properties of large collections of molecules, researchers have only recently managed to detect individual molecules. For example, attaching a highly fluorescent dye to a large protein molecule allows biophysicists to learn about details of the conformational changes that occur when the protein carries out its biological function. Now a team reports in the 14 December PRL a method for distinguishing the orientation of such a fluorescent molecule in a direct way. The technique may also be useful in understanding the structures of polymer matrices and other materials.
Fluorescence has been the method of choice for probing single molecules because fluorescence properties can be easily measured and are extremely sensitive to the immediate environment of the fluorescing molecule. In these experiments it is normally difficult to find the component of the electric dipole of the emitting molecule along the axis of the microscope which detects the fluorescence ( z-component), although it is possible to find the x and y components.
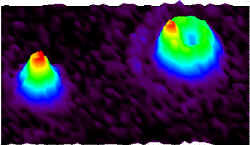
Robert Dickson, now of the Georgia Institute of Technology in Atlanta, was working on one approach to the problem when he realized that some of the image degradation in his system could be used to his advantage. “I came upon this by accident,” he recalls. “[The new method] is much simpler than the method I was going to use.” The technique Dickson and his colleagues developed relies on spherical aberration–the degradation of an image caused by light from different directions focusing at slightly different positions in the microscope. A fluorescent molecule aligned along the z-axis preferentially emits photons in the xy-plane, but some of them still enter the microscope at oblique angles. By contrast, a molecule aligned in the xy-plane sends many photons directly into the microscope lens. The team exaggerated the spherical aberration and caused z-aligned molecules to appear as out-of-focus doughnuts, with dark centers, while molecules in the xy-plane looked like bright disks.
Dickson and his colleagues embedded molecules of a dye in a thin layer of plastic on a piece of glass and illuminated it with laser light. By controlling the polarization of the laser they could selectively illuminate dye molecules in different orientations to show that the z-aligned molecules appeared as expected. They also ran computer simulations of the expected intensity patterns to further support their claims.
Although a few other research groups have measured the three-dimensional orientations of fluorescent molecules, those experiments relied on either indirect methods or very specialized setups. Dickson is most proud of the simplicity and directness of the new method, which should allow application in a wide variety of experiments, such as the interaction of two protein molecules, or as a probe of local conformations in a polymer matrix in which a dye can be embedded. Although he and his colleagues did not calculate the exact orientations of molecules in this paper, Dickson says that analysis of the intensity patterns, which he is now perfecting, leads directly to that information.
Jay Trautman, of SEQ Ltd. in Princeton, NJ, calls the work “elegant” and agrees that the method could be applied to experiments investigating the interactions of pairs of molecules. Shimon Weiss of Lawrence Berkeley National Laboratory says the ability to simultaneously measure orientations of many molecules in the microscope field-of-view “will have great utility for biophysical measurements of single macromolecules.”


