DNA Hairs Provide Potential for Molecular Self-Assembly
For many years, condensed-matter physicists have been involved with the fundamental problem of predicting how atoms and molecules behave collectively, exploiting their understanding for often unforeseen technological applications. More recently, the reconstruction of the atomic and molecular world on the nano- and microlength scales using human-engineered building blocks has come into focus. Nano- and microparticles of unconventional shape and surface properties have made their appearance in the laboratory and will hopefully become available in bulk quantities in the near future [1]. Particles can be “functionalized” by attaching specific molecules that change how areas of the particle respond to the surrounding solvent (e.g., creating hydrophobic/hydrophilic patches in the case of water), which allows some control over particle-particle interactions. With these tools, material scientists have a new challenge in designing particles that can self-assemble into novel materials for specific technological applications. A key part of this bottom-up strategy is understanding the effective potentials that control how the building blocks interact.
DNA-functionalized nanocolloids constitute a particularly promising class of new particles. In the simplest case, single strands of DNA are attached—in what is now a standard procedure—to the surface of a metal nanoparticle, completely shielding the metallic core and giving rise to a “hairy” particle. The choice of DNA as the functionalizing polymer is particularly rewarding. DNA is indeed highly selective in its interactions and is already exploited by nature in its role as genetic material. By choosing an appropriate sequence of bases, researchers can control the intra- and interparticle pairing abilities, achieving, in principle, a sophisticated and detailed handle on the (temperature-dependent) interparticle interaction potential. Additionally, in playing with single- or double-stranded DNA, scientists can tune the flexibility of the particle’s hairs.
Although the basic idea of coating colloids with DNA was presented more than a decade ago [2,3], it is only recently that experiments have reported the self-assembly of ordered crystalline phases [4,5]. Progress has been limited by several difficulties associated with the competing presence of kinetic traps, which favor the formation of amorphous aggregates rather than crystals [6]. One place where this poses a problem is in choosing the temperature for an experiment. Researchers must fine tune the temperature so that the system can reversibly sample a large fraction of microstates in order to find the ordered crystal structure. However, bath temperatures only a few degrees below the strand melting temperature are sufficient to turn the aggregation process into an irreversible one. The difficulties become worse if the desired crystal structure is something other than the most common cubic ones. In light of these complications, the community has long sought a way to accurately model the interparticle interaction potential, since that would be very valuable in predicting the most stable crystal structure as well as the optimal conditions for its formation.
To that end, Bianca Mladek of the University of Cambridge, UK, and University of Vienna, Austria, and co-workers report in Physical Review Letters on a promising model for predicting the phase diagram of DNA-functionalized nano colloids [7]. They focus on a binary mixture of nanocolloids that are functionalized with one of two different single-stranded DNA oligomers. The last bases of the two sequences are complementary, thus providing a binding site between distinct colloids.
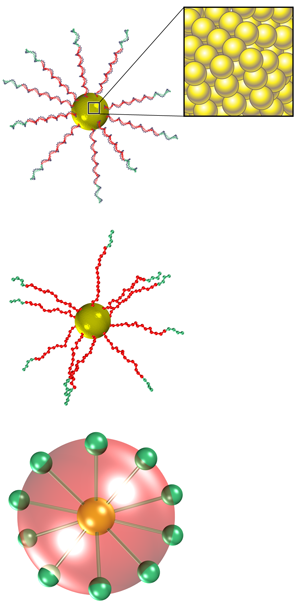
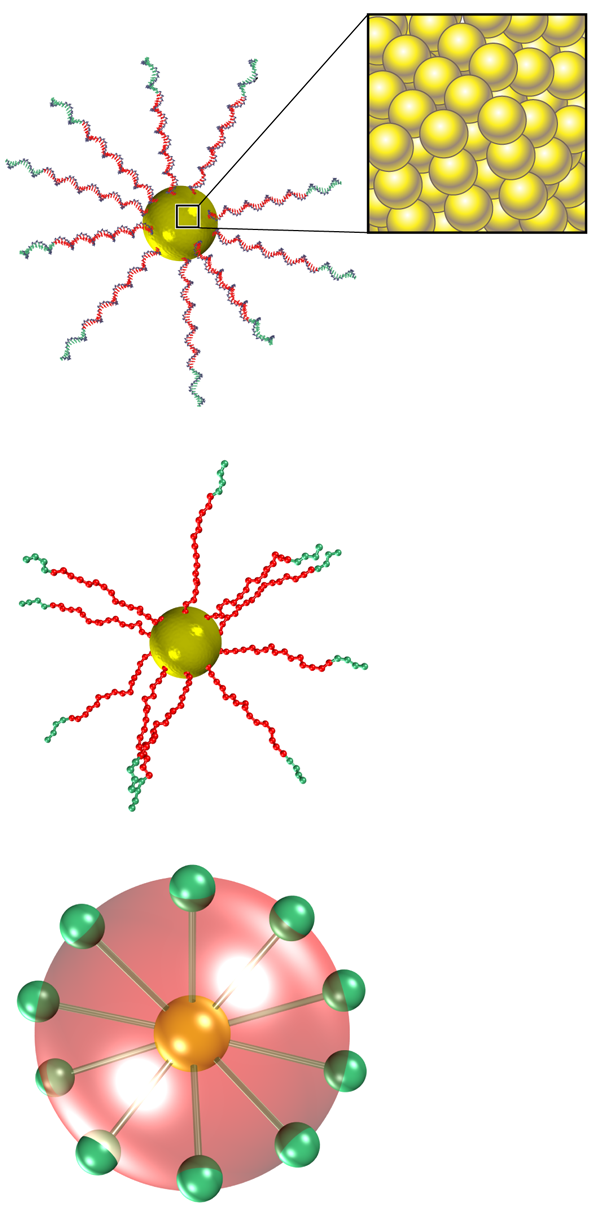
Mladek and co-workers faced two main obstacles in modeling the effective potential. The first was associated with the level of coarse-graining required to have a tractable interaction potential. For a single colloid, an atomistic simulation is, in principle, feasible, but the number of atomic degrees of freedom becomes intractable when investigating the collective behavior of multiple colloids. Instead, modelers must identify the important degrees of freedom and then apply a coarse-graining procedure that averages out the irrelevant ones. Mladek et al. employed a three-step coarse-graining procedure (see Fig. 1). First, they condensed all atoms defining a small group of nucleotide bases into a single representative interaction site. Second, they grouped all sites on the same DNA strand by whether they belonged or didn’t belong to the sticky binding sequence. As a result of these two steps, the degrees of freedom decreased to approximately per nanoparticle. The third and last step in the coarse-graining procedure involved condensing all remaining sites into two spherical components of the interaction potential, modeling the nonbinding and the binding contributions, respectively.
The second important problem to solve was the description of the hybridization binding free energy. Mladek and co-workers had to especially take care in preserving the specificity of the DNA-DNA interaction (the single-bond-per-strand—or valence preservation—condition [8]), which would have been lost in a standard coarse-graining process. This is a particularly important issue to take into account when particles interact via a lock-and-key type of interaction, in which paired strands become inert, i.e., unable to participate in subsequent bonds. The final binding probability that the authors used was parameterized against experimental data [9], providing the possibility of a close comparison between model predictions and laboratory results.
The multistep coarse-graining procedure provides a numerically tractable potential that makes it possible to evaluate the free energy of possible crystals and to calculate the range of temperatures and colloid concentrations over which the crystals are stable. For their modeled DNA nanocolloid, the researchers found that the most stable low-density ordered structure (which coexists with a dilute disordered fluid phase) is the same, mutatis mutandis, as the cesium-chloride ( ) crystal structure. The value of the lattice parameters, as well as their dependence on the external control parameters, properly describes the available experimental data, providing confidence in the effective potential stemming from the proposed coarse-graining process.
To better assess the generality of the model, further comparisons with additional experiments would be most welcome, e.g., with particles functionalized by DNA sequences differing in length and in coding sequence. The potential is here for an extremely powerful instrument that the soft matter and material science communities could use to optimize the properties of the nanoparticles by selecting the best size of the metal cores, the best DNA-grafting density, and the best DNA sequence. This will greatly facilitate the search for the optimal particles and for the best control parameters to make self-assembling crystals with particular targeted structures. With slight modification, the same approach can be adopted to help design and fabricate clusters using DNA-encoded nanoparticles [10].
References
- S. C. Glotzer and M. J. Solomon, “Dimensions in Anisotropy Space: Rationalizing Building Block Complexity for Assembly,” Nature Mater. 6, 557 (2007)
- C. A. Mirkin, R. L. Letsinger, R. C. Mucic, and J. J. Storhoff, “A DNA-based Method for Rationally Assembling Nanoparticles into Macroscopic Materials,” Nature 382, 607 (1996)
- A. P. Alivisatos, K. P. Johnsson, X. Peng, T. E. Wilson, C. J. Loweth, M. P. Bruchez, and P. G. Schultz, “Organization of Nanocrystal Molecules Using DNA,” Nature 382, 609 (1996)
- D. Nykypanchuk, M. M. Maye, D. van der Lelie, and O. Gang, “DNA-Guided Crystallization of Colloidal Nanoparticles,” Nature 451, 549 (2008)
- S. Y. Park, A. K. R. Lytton-Jean, B. Lee, S. Weigand, G. C. Schatz, and C. A. Mirkin, ”DNA-Programmable Nanoparticle Crystallization,” Nature 451, 553 (2008)
- S. Jahn, N. Geerts, and E. Eiser, “DNA-Mediated Two-Dimensional Colloidal Crystallization Above Different Attractive Surfaces,” Langmuir 26, 16921 (2010)
- B. M. Mladek, J. Fornleitner, F. J. Martinez-Veracoechea, A. Dawid, and D. Frenkel, ”Quantitative Prediction of the Phase Diagram of DNA-Functionalized Nanosized Colloids,” Phys. Rev. Lett. 108, 268301 (2012)
- J. Largo, P. Tartaglia, and F. Sciortino, “Effective Nonadditive Pair Potential for Lock-and-Key Interacting Particles: The Role of the Limited Valence,” Phys. Rev. E 76, 011402 (2007)
- N. R. Markham and M. Zuker, “DINAMelt Web Server for Nucleic Acid Melting Prediction,” Nucleic Acids Res. 33, 577 (2005)
- M. M. Maye, D. Nykypanchuk, M. Cuisinier, D. van der Lelie, and O. Gang, “Stepwise Surface Encoding for High-Throughput Assembly of Nanoclusters,” Nature Mater. 8, 388 (2009)