Maintaining the Sequence
Sequencing DNA is now routine, but reading out proteins is still challenging and expensive—it can take weeks and cost upwards of $70 per amino acid. Now Derek Stein and colleagues at Brown University, Rhode Island, have studied the feasibility of reading protein sequences by separating off amino acids one-by-one and spraying them into a mass spectrometer. There are many technical challenges that would have to be overcome to realize their method, but the researchers’ calculations indicate that it could work, enabling faster and cheaper sequencing methods for myriad proteins and polymers.
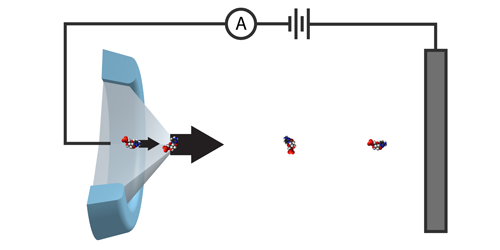
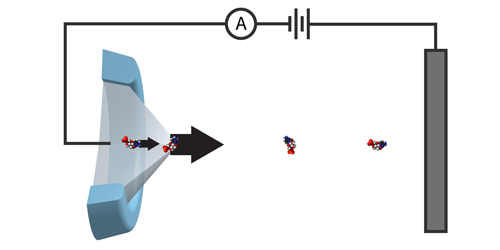
The group considered a row of connected particles—a protein made up of amino acids—traveling down a tube. At the exit of the tube, amino acids are cut off one-by-one, move a short distance, and are then identified by their mass and charge. For successful sequencing, the amino acids must retain their order until the readout process has been completed. The researchers predict this can occur if, as an amino acid is cut from the protein, it is simultaneously pulled away from it—something their calculations show could be done using an electrospray. The directional force imparted by the spray separates the particles, mitigating the effect of the Brownian motion that would otherwise jumble their order.
The team suggests two ways to cleave the amino acids: via laser light or using enzymes. For both methods, cutting has to occur within a narrow distance of the tube’s exit (3 nm for light and 100 nm for enzymes) to ensure the amino acids enter the mass spectrometer in sequence. Realizing this would require pinpoint cutting accuracies—something the researchers say is achievable with current technologies.
This research is published in Physical Review Applied.
–Katherine Wright
Katherine Wright is a Contributing Editor for Physics.