Low Efficiency Spotted in a Molecular Motor
Motor proteins are the engines of biology. They convert chemical energy into mechanical work to drive cell division, protein synthesis, muscle contraction, and other essential cellular processes. The motor protein kinesin carries vital cellular cargo through the cell by taking alternating steps along intracellular polymer tracks. A new study by Takayuki Ariga from Yamaguchi University, Japan, and colleagues has determined how much of the chemical energy is converted to mechanical work and how much is lost to heat for one type of kinesin called kinesin-1 [1]. The experiments, which use optical tweezers to exert an oscillating force on single motor proteins, show that kinesin-1 loses about 80% of its input energy to dissipation, or heat, within the molecule as it moves. Understanding the energy conversion by kinesin-1 compared to other motor proteins that are much more efficient could lead to insight into how the mechanochemistry of motor proteins is tuned for their specific cellular roles.
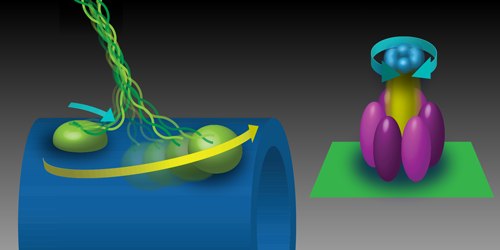
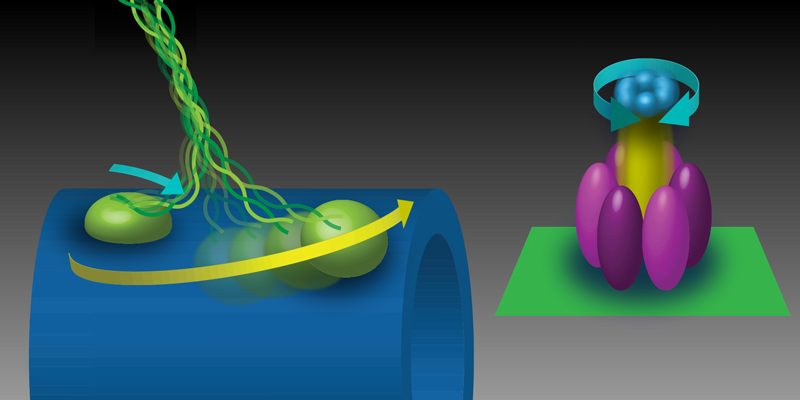
Kinesin-1 plays a central role in intracellular transport, carrying vesicles, organelles, and signaling molecules outward from the cell center [2]. The tail domains of kinesin attach to its cargo, while the two motor domains on the other end bind to microtubules, which are the polymer tracks that form part of the cytoskeleton (Fig. 1). Stepping is powered by the fuel molecule ATP (adenosine triphosphate). When an ATP molecule binds to kinesin, the ATP breaks up into ADP (adenosine diphosphate) and an inorganic phosphate. This reaction, called ATP hydrolysis, releases about 85 pN nm (or ∼10−19J) of energy, which is used to drive a structural change in one of the motor domains that causes the other domain to move towards the next binding site on the microtubule. Previous work has shown that kinesin-1 takes hundreds of these steps along a microtubule, traveling at roughly 1𝜇m∕s. However, mutations in kinesin-1 or its binding partners can disrupt transport, leading, in some cases, to neurodegenerative disease [3].
Much of our understanding about kinesin-1 and related motor proteins was made possible by revolutionary techniques that allow researchers to observe and manipulate single molecules. One technique—optical trapping—has been particularly useful, as evidenced by half of the 2018 Nobel Prize in Physics being awarded to its developer, Arthur Ashkin (see 4 October 2018 Focus story). Optical traps or tweezers use a tightly focused laser beam to manipulate micrometer-sized objects with piconewton-level forces. By attaching kinesin motors to a latex bead and then holding the bead in an optical trap, researchers have measured the motor’s step size (8 nm) and the maximum force, or load, that the motor can carry forward (6 pN). While the first of these measurements was reported over 20 years ago [4], optical trapping assays continue to provide insights into kinesin-1 and other motor proteins [5].
A rough estimate of kinesin’s energy efficiency assumes that one ATP molecule is hydrolyzed for each 8-nm step. At kinesin-1’s maximum load of 6 pN, the resulting efficiency would be ∼60%. However, this estimate does not account for futile ATP hydrolysis due to the frequent backward steps that kinesin-1 undergoes at high loads. To obtain an accurate measurement, Ariga and coauthors performed optical tweezer measurements at low loads where backward steps are rare. They applied an oscillating force to a single kinesin-1 molecule and tracked its step motion along a microtubule. By analyzing the fluctuations in the motor’s velocity, the team was able to quantify both the equilibrium dissipation due to viscous forces and the nonequilibrium energy losses within the kinesin molecule itself. When they compared the work and dissipation to the total free energy available from ATP hydrolysis, they found that only 20% of the free energy was accounted for, indicating that 80% of the energy is lost to internal dissipation (or heating) of the kinesin molecule as it walks.
These observations are particularly striking when compared to another motor protein, the F1 subunit of ATP synthase (F1-ATPase). This rotational motor synthesizes ATP from ADP and inorganic phosphate. A previous analysis of F1-ATPase concluded that it converts close to 100% of the available energy to mechanical work [6]. What differences in the mechanochemistry of these two motors explain their disparate efficiencies? For kinesin, Ariga et al. show that its low efficiency is not caused by futile ATP hydrolysis due to backward steps nor by energy lost to stretching the kinesin-1 molecule. They conclude that the inefficiency of kinesin is likely linked to its irreversibility. When a kinesin steps backwards, it does not create energy by generating ATP fuel [7]. By contrast, the motion of F1-ATPase is a reversible process. In the “forward” direction, it hydrolyzes ATP to rotate its shaft counterclockwise. However, if turned in the “backward” direction, it can generate ATP from ADP and inorganic phosphate.
Comparisons with other motor proteins may offer alternative explanations for kinesin-1’s low efficiency. For example, optical trapping measurements on a closely related kinesin, kinesin-8, indicate that kinesin-8 experiences friction between the motor and the microtubule track as it steps [8]. This friction arises from the continuous formation and breakage of bonds. If kinesin-1 experiences similar friction during stepping, the corresponding dissipation would account for 5–80% of the total energy available. Thus, motors that translocate along a track might pay an energetic penalty to maintain their association with the track. In contrast, rotary motors like F1-ATPase might maximize efficiency by rotating about a nearly frictionless bearing [9].
These experiments highlight how the mechanochemistry of different motor proteins is adapted for their cellular roles. Further, they provide a framework for comparing motor proteins based on their energetics. There are more than 45 kinesins, with roles in intracellular transport, regulation of microtubule polymerization, and organization of the microtubule cytoskeleton. These motors vary widely with respect to velocity, processivity, and force generation. Comparing motors with different properties, there is an opportunity to uncover how motors are tuned for their specific functions by linking energetics with motor mechanochemistry. These principles can be used to help us understand how these tiny biological engines may have evolved and how defects in their function can lead to disease [3]. Engineers may also apply these functional analyses to the design of nanodevices that mimic or utilize motor proteins [10].
This research is published in Physical Review Letters.
References
- T. Ariga, M. Tomishige, and D. Mizuno, “Nonequilibrium energetics of molecular motor kinesin,” Phys. Rev. Lett. 121, 218101 (2018).
- R. D. Vale, T. S. Reese, and M. P. Sheetz, “Molecular motors in neurons: transport mechanisms and roles in brain function, development, and disease,” Cell 42, 39 (1985).
- N. Hirokawa, S. Niwa, and Y. Tanaka, “Molecular motors in neurons: Transport mechanisms and roles in brain function, development, and disease,” Neuron 68, 610 (2010).
- K. Svoboda, C. F. Schmidt, B. J. Schnapp, and S. M. Block, “Direct observation of kinesin stepping by optical trapping interferometry,” Nature 365, 721 (1993).
- M. Capitanio and F. S. Pavone, “Interrogating biology with force: Single molecule high-resolution measurements with optical tweezers,” Biophys J. 105, 1293 (2013); F. M. Fazal and S. M. Block, “Optical tweezers study life under tension,” Nat. Photon. 5, 318 (2011).
- S. Toyabe, T. Watanabe-Nakayama, T. Okamoto, S. Kudo, and E. Muneyuki, “Thermodynamic efficiency and mechanochemical coupling of F1-ATPase,” Proc. Natl. Acad. Sci. U.S.A. 108, 17951 (2011).
- B. E. Clancy, W. M. Behnke-Parks, J. O. L. Andreasson, S. S. Rosenfeld, and S. M. Block, “A universal pathway for kinesin stepping,” Nat. Struct. Mol. Biol. 18, 1020 (2011).
- V. Bormuth, V. Varga, J. Howard, and E. Schaffer, “Protein friction limits diffusive and directed movements of kinesin motors on microtubules,” Science 325, 870 (2009).
- K. Okazaki and G. Hummer, “Elasticity, friction, and pathway of 𝛾-subunit rotation in F0F1-ATP synthase,” Proc. Natl. Acad. Sci. U. S. A. 112, 10720 (2015).
- H. Hess and G. Saper, “Engineering with biomolecular motors,” Acc. Chem. Res. (2018).