Brain Motion Under Impact
Strike a bell or a drum and its sound will be dominated by a few vibrational patterns (“modes”), each of which has a characteristic shape, frequency, and damping rate. A new study has found that the mechanical response of the human brain to impact can similarly be described with a small number of dynamic modes. Kaveh Laksari of the University of Arizona in Tucson, Mehmet Kurt of the Stevens Institute of Technology in New Jersey, and colleagues used measurements of head impacts during football games as input to a computer model that simulates the point-by-point displacement of tissue within the brain [1]. Their analysis of the simulated motion shows that this “displacement field” can be described with just a few modes whose characteristic frequencies are around 20 to 40 Hz. Their findings could offer insight into the mechanics of traumatic brain injury (TBI), perhaps leading to new designs for protective equipment.
TBI is a pervasive and serious medical problem. “Mild” TBI, otherwise known as a concussion, is common in sports [2]; more severe injuries arise from automobile accidents, falls, or military combat [3]. Even impacts that don’t rise to the level of a concussion can, when repeated many times, cause a neurodegenerative condition known as chronic traumatic encephalopathy [4]. This is why researchers have grown increasingly concerned about athletes playing contact sports such as football.
The general mechanical principles behind most TBIs are well understood. During an impact, the skull accelerates rapidly. The brain, which has considerable inertia and is surrounded by cerebrospinal fluid, accelerates slightly less quickly than the skull and deforms under forces from the various membranes, blood vessels, and connective tissue that attach it to the skull. This rapid deformation, or strain, of brain tissue stretches and injures delicate nerve fibers, degrading brain function.
This general picture is not, however, sufficient for predicting the short- and long-term effects of TBI. And, unfortunately, the more detailed mechanics of head impacts remain largely mysterious. That’s partly because brain deformation is challenging to observe: The brain is well hidden and any attempt to expose it could change its behavior or disrupt its structure and functioning. Another issue is the complexity of the brain as a material: It is both heterogeneous (different locations in the brain have different material properties) and anisotropic (it responds differently to forces applied along different directions) [5]. Also, brain tissue’s response to a deforming force isn’t perfectly spring-like but is instead nonlinear and damped [6]. Finally, every head is unique, and each impact differs in location, magnitude, and direction. This combination of features makes it difficult to say precisely how the brain deforms during either mild or severe TBI.
Researchers have set out to gather this missing information in various ways. On the experimental front, some groups have used magnetic resonance imaging (MRI) [7, 8] to measure brain motion in volunteers whose heads were subjected to reasonably mild (not injury-producing) accelerations. Other groups have measured brain motion at injury-level accelerations in the brains of cadavers [9]. An alternative tack is to rely on detailed finite-element computer simulations [10, 11], which simultaneously solve equations that describe small, individual elements of brain tissue. This approach can predict brain motion for a given head acceleration, using different material models to describe the skull, brain, cerebrospinal fluid, and membranes.
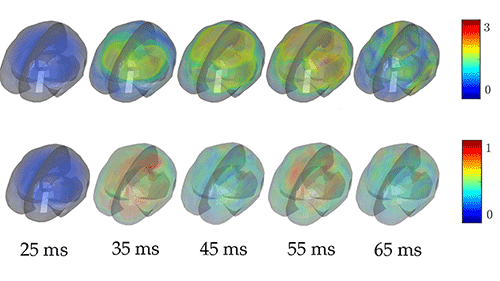
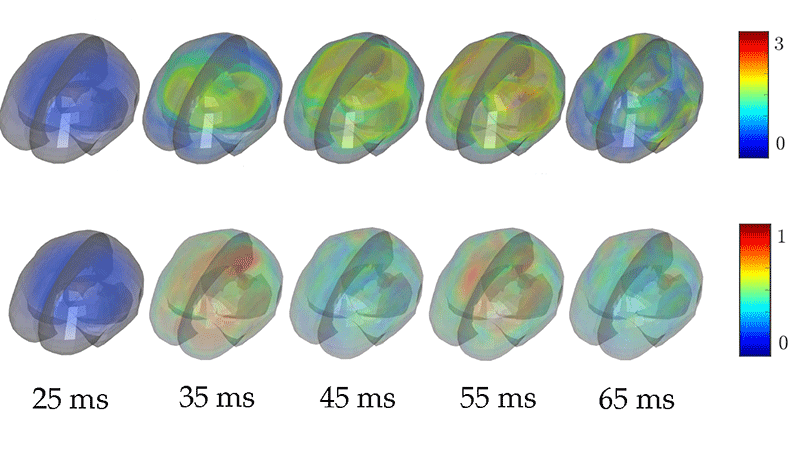
Laksari et al. [1] have added a new twist to the classic simulation approach by analyzing simulated brain motion with a technique called dynamic mode decomposition (DMD). This technique is roughly analogous to a Fourier analysis, which describes a signal in terms of a set of predefined functions (say, sines and cosines). The difference is that the modes in DMD are obtained anew from each set of data (simulated or measured), which allows the data to be represented with fewer modes. This data-driven approach is effective for describing the displacement field in a bounded, heterogeneous material like the brain, particularly during its damped transient motion after a head impact. The researchers used a detailed finite-element model [10] to simulate the displacement and strain fields from head impacts like those that football players incur during a game. As an input to this model, they used 187 measurements of head accelerations in players, which they had obtained previously from sensor arrays mounted in the players’ mouth guards. Using DMD, they broke down the simulated fields into a superposition of a few damped, oscillating modes. They then determined which modes contributed the most energy.
Strikingly, the authors found that they could closely approximate the displacement and strain fields using only a few low-frequency ( <40 Hz) modes (Fig. 1). Moreover, the strains from mild impacts, they showed, could be accurately reproduced with a single mode, while a larger number of modes was required to model more severe impacts. This difference makes sense because structural and material nonlinearities play a greater role in severe impacts, creating more complex strain fields.
The team’s findings have at least two interesting implications. First, researchers will potentially be able to model and simulate TBI in football more efficiently, since only a small number of modes are needed to describe the impact. Second, the findings suggest that football helmets and other protective gear should be designed to prevent the excitation of low-frequency modes, rather than to minimize the peak acceleration that the head experiences. In fact, researchers have long been puzzled by the lack of correlation between peak acceleration and injury. This study illustrates why such a relationship may not actually exist.
As Laksari et al. note, however, there is a caveat: their conclusions are drawn from simulations that have not yet been systematically compared to MRI measurements on living humans. The team has compared the displacements in their simulations to those obtained from cadaver experiments. But this comparison does not establish the accuracy of their predictions because a cadaver brain only approximately corresponds to the brain in a living human. Fortunately, relevant data from MRI studies are available and a comprehensive test of the new predictions should soon be possible. The work from Laksari et al. may, in fact, resolve another problem, which is how best to quantitatively compare simulated data with MRI measurements. Their study suggests that the frequencies, damping rates, and spatial features of the relevant dynamic modes are excellent metrics for such comparisons.
This research is published in Physical Review Letters.
References
- K. Laksari, M. Kurt, B. Hessam, S. Kleiven, and D. Camarillo, “Mechanistic Insights into Human Brain Impact Dynamics through Modal Analysis,” Phys. Rev. Lett. 120, 138101 (2018).
- W. P. Meehan III and R. Mannix, “Pediatric Concussions in United States Emergency Departments in the Years 2002 to 2006,” J. Pediatr. 157, 889 (2010).
- C. W. Hoge, D. McGurk, J. L. Thomas, A. L. Cox, C. C. Engel, and C. A. Castro, “Mild Traumatic Brain Injury in U.S. Soldiers Returning from Iraq,” N. Engl. J. Med. 358, 453 (2008).
- A. C. McKee et al., “The Spectrum of Disease in Chronic Traumatic Encephalopathy,” Brain 136, 43 (2012).
- J. L. Schmidt, D. J. Tweten, A. A. Badachhape, A. J. Reiter, R. J. Okamoto, J. R. Garbow, and P. V. Bayly, “Measurement of Anisotropic Mechanical Properties in Porcine Brain White Matter Ex Vivo Using Magnetic Resonance Elastography,” J. Mech. Behav. Biomed. Mater. 79, 30 (2018).
- F. Velardi, F. Fraternali, and M. Angelillo, “Anisotropic Constitutive Equations and Experimental Tensile Behavior of Brain Tissue,” Biomech. Model. Mechanobiol. 5, 53 (2005).
- A. A. Sabet, E. Christoforou, B. Zatlin, G. M. Genin, and P. V. Bayly, “Deformation of the Human Brain Induced by Mild Angular Head Acceleration,” J. Biomech. 41, 307 (2008).
- A. K. Knutsen, E. Magrath, J. E. McEntee, F. Xing, J. L. Prince, P. V. Bayly, J. A. Butman, and D. L. Pham, “Improved Measurement of Brain Deformation During Mild Head Acceleration Using a Novel Tagged MRI Sequence,” J. Biomech. 47, 3475 (2014).
- W. N. Hardy, C. D. Foster, M. J. Mason, K. H. Yang, A. I. King, and S. Tashman, “Investigation of Head Injury Mechanisms Using Neutral Density Technology and High-Speed Biplanar X-Ray,” Stapp Car Crash J. 45, 337 (2001).
- S. Kleiven, “Predictors for Traumatic Brain Injuries Evaluated through Accident Reconstructions,” Stapp Car Crash J. 51, 81 (2007).
- S. Ji, W. Zhao, Z. Li, and T. W. McAllister, “Head Impact Accelerations for Brain Strain-Related Responses in Contact Sports: A Model-Based Investigation,” Biomech. Model. Mechanobiol. 13, 1121 (2014).