First Frame from 3D Molecular Movie
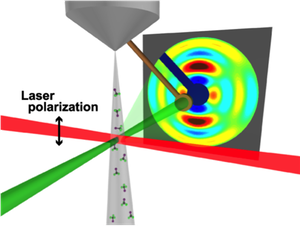
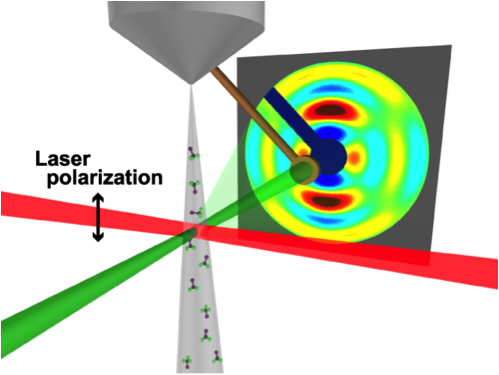
Still images of molecules are not new, but a highly anticipated “coming attraction” is a movie showing fast changes in a molecule’s structure. Research reported in Physical Review Letters brings this film closer to reality with the first 3D shot of a molecule in a gas. The authors aligned their gas molecules with a laser pulse and then hit them with a very short blast of electrons. The pattern of the scattering electrons provided enough information to make an image. If several of these images can be strung together in succession, then researchers could view the rapid shape changes in biological molecules, such as retinal, which is involved in vision, or chlorophyll, which is central to photosynthesis.
The standard way to measure a molecule’s structure is to scatter x rays off a crystalline sample and use the pattern of x-ray spots (the diffraction pattern) to work back to the structure. However, researchers sometime can’t—or may not want to—lock a molecule into a crystal. To study free molecules in a gas, they have turned to electrons, which provide higher resolution than x rays and also allow for a lower beam intensity. But in gas-electron diffraction, the molecules are randomly oriented, so the diffraction pattern is a blurry average over all orientations. Building a 3D picture requires an initial guess about where the atoms are. “This starts to pose a problem when you work with very large molecules,” says Martin Centurion of the University of Nebraska, Lincoln.
One solution researchers have used is to align the molecules with the polarization (electric field) direction of a laser that shines on the gas at the same time as the electron beam [1]. However, the laser light not only aligns but also deforms the molecules, so you don’t see them in their natural state. It’s also difficult to obtain both perfect alignment and a strong diffraction signal. Centurion and his colleagues have a new technique that addresses these challenges by using, first, a laser pulse that generates alignment some time after the pulse ends, and, second, an algorithm that can correct the data for imperfect alignment.
For their first demonstration, the team chose a highly symmetric molecule, trifluoroiodomethane ( CF3I). Previous work has shown that the carbon-iodine bond in this molecule forms a central axis, with the three fluorine atoms extending out from the carbon like a tripod (similar to methane, CH4). The team exposed a gas jet of CF3I molecules to a 300-femtosecond laser pulse, which essentially induced a molecular rotation rate that, for each molecule, depended on the degree to which it was initially misaligned with the laser’s polarization direction. The maximum alignment of C-I axes occurred a few picoseconds after the laser was off. At this moment, the team hit the sample with a 500-femtosecond electron-beam pulse and recorded the diffraction pattern—the first of its kind with subpicosecond resolution.
To acquire enough information for the 3D structure, the researchers collected diffraction patterns for two alignment directions ( 90 degrees and 60 degrees, relative to the electron beam) and one pattern with no laser alignment. The team’s new computer algorithm extracted from the data a structure whose interatomic distances and bond angles were in agreement with previous observations by other techniques.
The new technique, in principle, allows 3D imaging of large molecules that can’t be crystallized, but the team is focusing on movies of molecules whose fast structural changes are unknown. With some improvements to their experiment, such as a shorter electron pulse, the first molecular movies could be coming out in a year or so, Centurion says. One of the first “actors” might be retinal, a visual-system molecule that rapidly changes shape when it absorbs light. “We know what it looks like before and after, but not in between,” Centurion says.
“Although the authors have produced only a single frame of a hypothetical 3D movie that shows molecules evolving over time, they have clearly demonstrated the basic principle,” says Allen Landers of Auburn University in Alabama. But he sees many challenges ahead, especially as the complexity of the target molecule increases. Vincent McKoy of the California Institute of Technology in Pasadena points out that less symmetric molecules will need to be aligned on two axes, not just one, which will be challenging. “At present this looks like a promising method for structural determination, but the road from here to movies might be long,” he says.
–Michael Schirber
Michael Schirber is a Corresponding Editor for Physics Magazine based in Lyon, France.
References
- K. Hoshina, K. Yamanouchi, T. Ohshima, Y. Ose, and H. Todokoro, “Alignment of CS2 in Intense Nanosecond Laser Fields Probed by Pulsed Gas Electron Diffraction,” J. Chem. Phys. 118, 6211 (2003)
More Information
description of gas electron diffraction by Derek Wann of the University of Edinburgh