Entropy Gives DNA a Shove
DNA isn’t just for cloning sheep. The stringy stuff also ideally illustrates the effects of entropy, the elusive thermodynamic property that reflects disorder. In the 25 March print issue of PRL, physicists have demonstrated its newest entropic trick: an elongated DNA molecule squirts from a confined space into an open one under the propulsion of entropy alone. The trick may translate into chip technology for sorting DNAs by size, enabling fast DNA fingerprinting at airports.
In DNA and other polymers, in which tens or even millions of smaller molecules link in a chain, the amount of entropy depends on the freedom of the chain to swivel about each link. Each free link on a chain multiplies the molecule’s potential shapes and elevates the entropy, according to a theory from the 1930s. And like any other system, a fixed length of DNA cannot resist a chance to increase its entropy.
It might seem strange that the availability of options could exert a force, but that’s what tugs against you whenever you stretch a rubber band. The resistance mounts as the stretch aligns and constrains the shapes of polymers in the rubber. DNA is elastic too, because it likewise loses entropy when stretched. Theorists have supposed that an entropic resistance to unraveling is also what slows DNAs that worm through the microscopic meshwork of watery gels–as electric fields compel them to do every day on laboratory bench tops around the world. Yet the entropic force of such confinement isn’t so easy to isolate or measure.
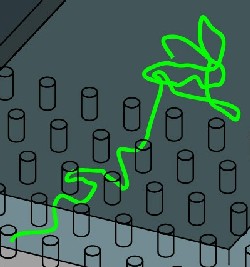
Enter Steve Turner and collaborators at Cornell University in Ithaca, NY. Taking a technique from the semiconductor industry, they etched a shallow trough into a chip and obstructed it with a forest of tiny pillars at the midpoint. When they filled the trough with water and added large pieces of fluorescently stained DNAs, the team could watch the motions of single molecules through a microscope. Outside the pillar forest, the DNAs could adopt only garden hose-like arrangements in two dimensions. To penetrate the crowded pillars, they had to straighten out.
Applying a voltage drew the negatively charged DNAs rapidly into the forest–at the expense of entropy. Shutting off the voltage caused any molecule with a free end to squirt back out again, as a videotape revealed. The molecules, which took tens of seconds to eject, gained speed as they exited like chains sliding off a table.
The forest barrier suggests a method for sorting DNAs by size, Turner says. Brief voltage pulses should embed short molecules fully into the forest with the first blow and kick them across with subsequent ones. Longer molecules would poke in just partially before each pulse ended, only to bounce back without making headway. Because enzymes chop different people’s genomes differently, Turner points out, sorting DNAs by size can produce a fingerprint.
Others just find the experiment nifty. “This is really a graphic illustration that things are behaving as we supposed,” says David Hoagland of the University of Massachusetts at Amherst. In particular, he says, DNA’s accelerating extrusion confirms that each emerging segment adds equally to the entropy of the chain. But other candidates exist for sorting DNA on a chip for airports and crime scenes. Only more development and testing will decide the winner, he says.
–Oliver Baker
Oliver Baker is a freelance science writer based in Davis, CA.


