Identifying Cellular Concentrations via Fluctuations
Tissues are complex systems formed of a variety of different cells with different functions and genetically driven biochemistry. It is very rare that biological samples contain only one cell type. The question then arises, is it possible to measure samples and calculate exactly what cells are present, how many of each cell there are, and which genes they each individually express?
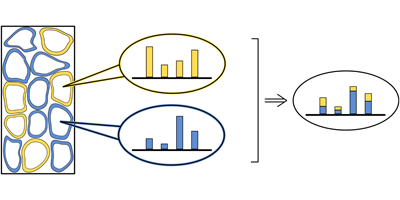
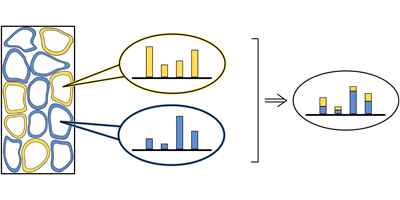
The answer is yes, but only averaged values of the different cell types in a given tissue can be obtained experimentally. Writing in Physical Review E, Nico Riedel and Johannes Berg from the University of Cologne, Germany, show that statistical mechanics can be used to solve this problem. The pair start with an artificial computational tissue from which they take a number of different samples. Due to fluctuations, the samples contain slightly different proportions of each cell type, which result in sample-to-sample variations of the gene expression levels. To reconstruct the composition of the tissue and gene expression of the component cells, Riedel and Berg designed a deconvolution algorithm, which exploits these sample-to-sample fluctuations.
The algorithm developed by Riedel and Berg can require as few as five different samples to accurately determine the tissue composition, making this an experimentally attractive method. However, the method has only been tested on artificial datasets, so it remains to be seen whether it can be accurately applied to real tissues. If so, this technique has wide-ranging medical applications and could be used to determine the proportion of healthy and abnormal cells in tissues and tumours, or the composition of cells in blood with only relatively small samples being needed. – Katherine Thomas