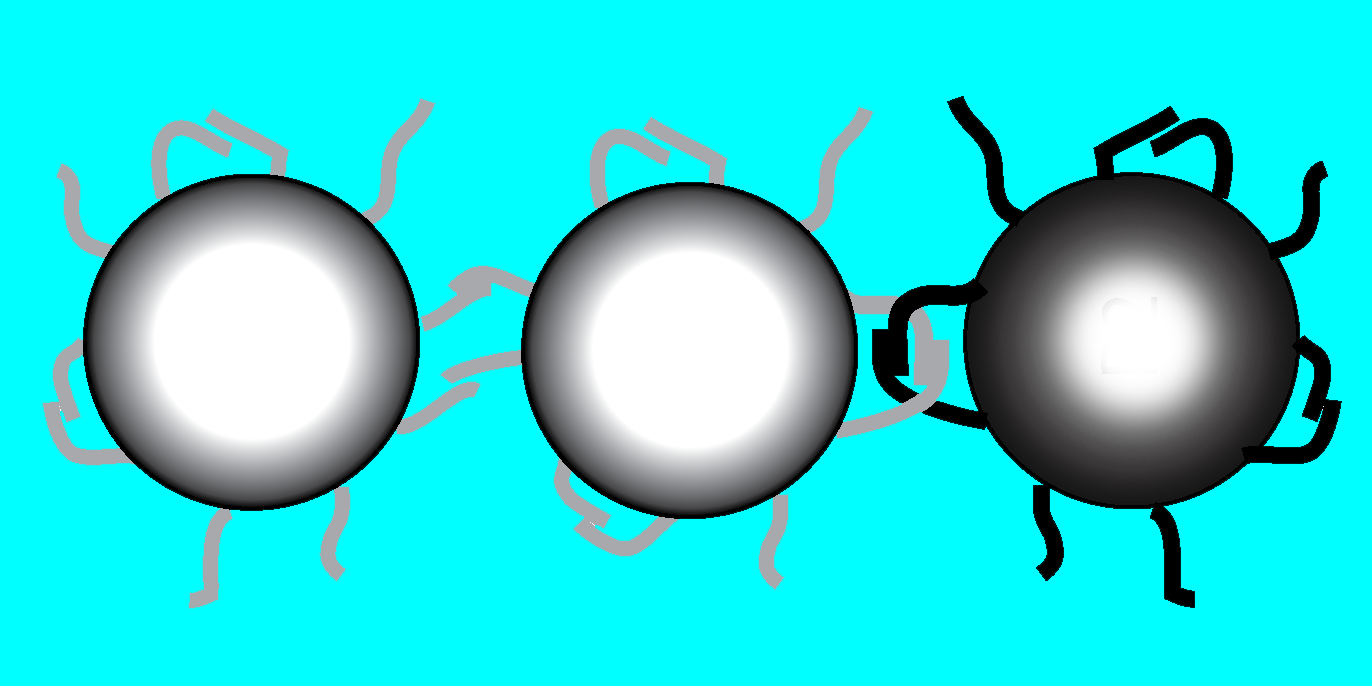
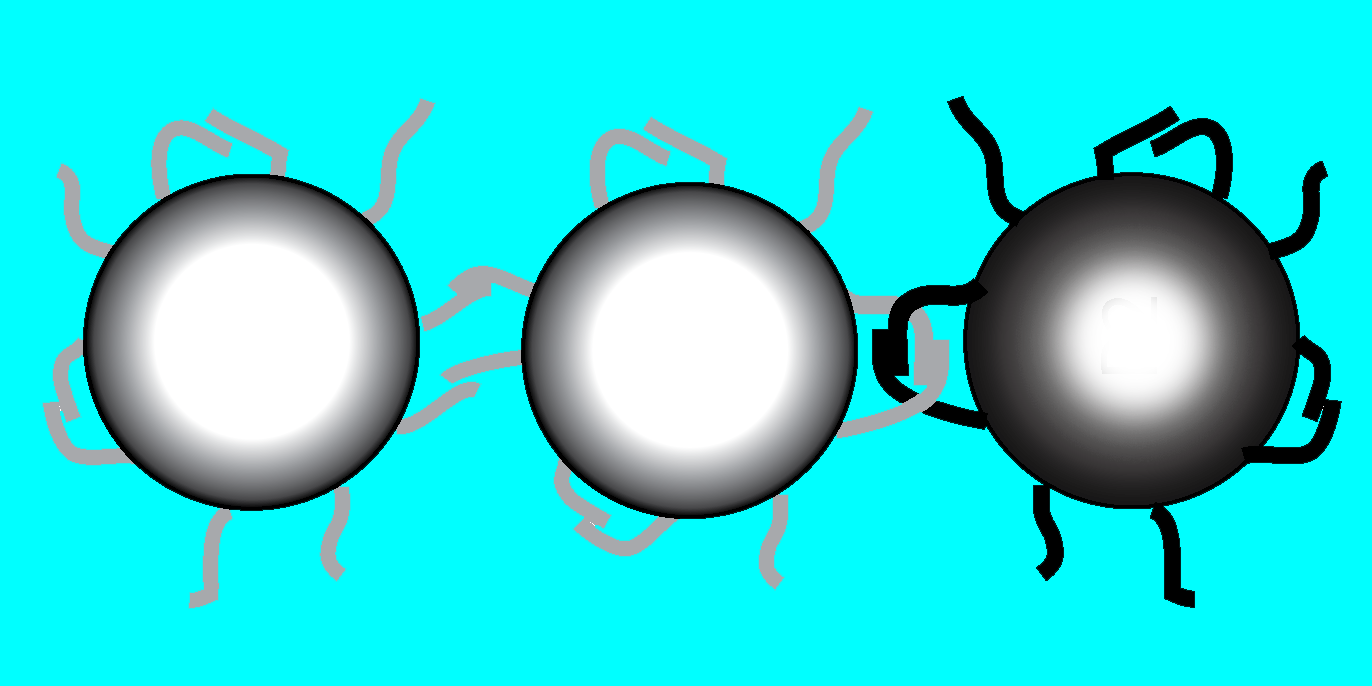
Molecular Velcro
Surprisingly complex structures arise from tailoring the interactions between tiny building blocks and letting the blocks self-assemble. One powerful way to select which blocks stick together is to adorn them with engineered DNA that binds only to other blocks with DNA having a complementary sequence. In Physical Review Letters, Lang Feng and colleagues from New York University demonstrate a different way to stick tiny particles together, by entangling loops of DNA on neighboring particles.
To promote loops, the team decorated one-micron-diameter polystyrene spheres with many DNA molecules having a self-complementary sequence. Such molecules can each bind to their own mirror-image sequence in another molecule nearby. They then mixed together spheres adorned with two incompatible sequences. A neighboring pair of spheres with the same sequence could bind to each other directly, but a pair with different sequences could only form loops on each sphere separately. Nonetheless, the spheres will bind if the loops intertwine.
The team used spheres containing a magnetic core, which join up in chains when a small magnetic field is present. This ensured that the spheres were close together as the loops formed, which happened when the researchers cooled the mixture. Even after they removed the magnetic field, many spheres remained bound. But adding an enzyme that lets DNA strands cross each other intact broke up the chains, as expected for entangled loops. Unlike traditional DNA binding, this Velcro®-like attachment depends on the order of processing steps, and could provide new ways to assemble structures. – Don Monroe