Through the Eye of the Needle
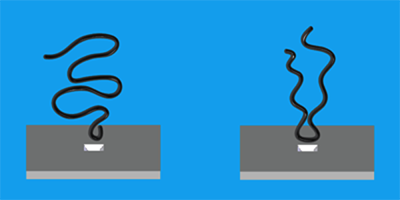
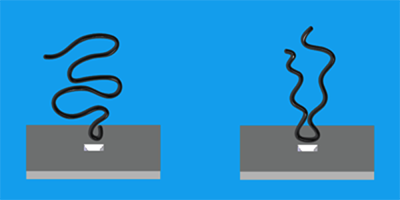
Nanopores are nanometer-sized holes in membranes that show promise as DNA sequencers: researchers can monitor the opening and closing of a pore as chains of nucleic acids pass through, as if they were tugging on a strand of pearls. Although typically done with single unfolded biomolecules, larger nanopores can also pull in more than one strand, as might occur with a folded molecule. In Physical Review Letters, Mirna Mihovilovic and colleagues at Brown University, Rhode Island, report experiments that should help clarify how different molecular configurations are captured and move through a nanopore. In particular, they observe that DNA has a favorable tendency to enter the pore end-first.
The authors fabricated a pore nm in diameter through a sheet of silicon nitride separating two conducting aqueous solutions. An applied voltage generates a current that varies according to how much the pore is blocked. The time course of the ionic current indicated whether single strands, multiple strands, or multiple folds of the same strand were passing through, or whether the molecule was a fragment or a damaged strand. From this, Mihovilovic et al. could amass data on capture as a function of the location at which the strand is trapped.
The Brown team found that the probability for a strand to be pulled into the pore at a particular location increased rapidly close to the ends of the polymer. Previously, researchers had assumed that the likelihood of capture was constant along the chain. Their modeling shows that this is due to maximization of configurational entropy: there are many more ways for a polymer to approach the pore end-first than leading with a segment near the middle. – David Voss