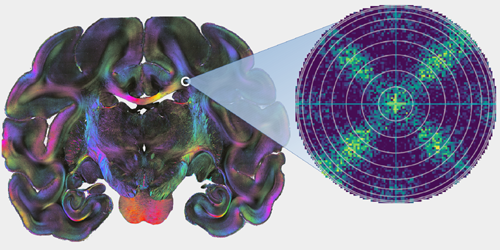
Untangling Neurons with Scattered Light
To understand how function emerges from the brain’s structure, neuroscientists need to know the intricate three-dimensional pathways followed by nerve fibers. State-of-the-art polarization microscopy techniques reveal 3D fiber pathways with micrometer resolution but cannot reliably determine fiber crossing points. Now, Miriam Menzel, at Jülich Research Center in Germany, and colleagues show how light scattering can help to detect fiber crossings in those 3D images, revealing the network’s substructure. The method allows a more detailed network model of the brain and may reveal tissue changes responsible for neurodegenerative diseases.
Three-dimensional polarization microscopy maps nerve fiber networks in hair-thin brain sections by measuring the 3D nerve fiber orientation in each micrometer-scale tissue voxel. When a voxel contains two fibers crossing in the same plane, the technique often returns a single, spurious orientation, misinterpreting the fibers as pointing out of the section plane. To overcome this limitation, Menzel and colleagues sought additional information using conventional transmission microscopy. In brain sections with known fiber configurations, they demonstrated that the transmitted light intensity depends on the angle between the fibers and the direction of light propagation. Using numerical simulations of polarization microscopy in fibrous brain tissue, the researchers found that this extra information can distinguish crossing in-plane fibers from fibers that point out-of-plane. In further microscopy studies, they showed that scattered light measurements allowed them to reconstruct the brain’s tissue substructure with unprecedented detail, including the angles at which nerve fibers cross.
The improved reconstruction of nerve fiber pathways, especially fiber crossings, could lead to better interpretation of medical scans, such as diffusion MRI. The simulation framework and results could also be generalized to other microscopy techniques and fibrous tissue samples.
This research is published in Physical Review X.
–Rachel Berkowitz
Rachel Berkowitz is a Corresponding Editor for Physics based in Seattle, Washington, and Vancouver, Canada.