Distinguishing Noise Sources with Information Theory
From neurons firing in the brain to chromosomes moving in a nucleus, biology is full of systems that can be modeled as a network of interacting particles in a noisy environment. But fundamental questions remain about how to reconstruct the particle interactions from experimental observations [1] and about how to separate the noise coming from the environment from the intrinsic interactions of the system [2]. Now, Giorgio Nicoletti of the University of Padua, Italy, and Daniel Busiello of the Swiss Federal Institute of Technology in Lausanne show how “mutual information” can help answer this second question [3]. Their theory disentangles the roles of the stochastic environment and the deterministic interaction forces of a system in creating correlations between the positions of two particles. This step is an important one toward understanding the role of environmental noise in real systems, where that noise can produce a response as complex as the signal produced by the system’s interactions.
Thinking back to my high school biology lessons, biological processes were always presented as deterministic, meaning there is no randomness in how they progress. For example, I was taught that a molecular motor marches rhythmically down a tubule, while ions traverse a membrane at regular intervals. But these processes are in fact inherently random and noisy: A molecular motor’s ‘‘foot’’ diffuses around before taking a step, and the motor can completely detach from the tubule; ion gates fluctuate between ‘‘open’’ and ‘‘closed’’ at random time intervals, making the flow of ions intermittent. This issue has led me and other researchers to wonder how these systems can properly function given the constant presence of fluctuations, or even more surprisingly, how these systems can gain improved function through these fluctuations.
Roughly speaking, the fluctuations and interactions associated with a system can be categorized as either “intrinsic” or “extrinsic,” where intrinsic fluctuations and interactions are typically those caused by the system itself (e.g., the random opening and closing of ion gates) and extrinsic fluctuations and interactions arise outside the system (e.g., thermal changes in the environment in which the ions diffuse or fluctuations in the ion-gate-activation signals). Extrinsic fluctuations can mask signals related to subtle internal changes of a system and add correlations that resemble those that arise from intrinsic interactions. Both factors make it more difficult to extract model parameters or even interaction laws. Nicoletti and Busiello address this issue by using mutual information theory, which quantifies the level of independence of two variables, to untangle internal and environmental correlations from a signal.
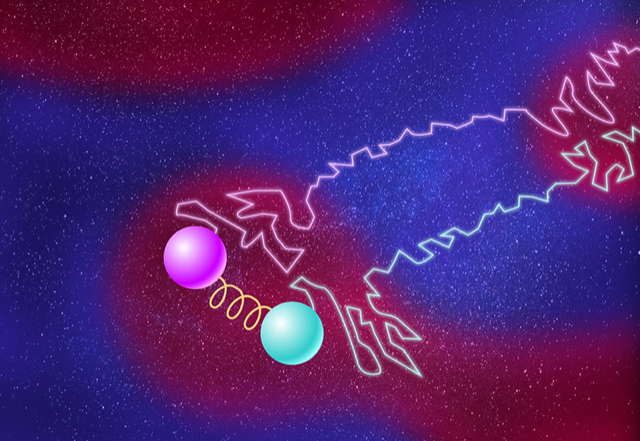
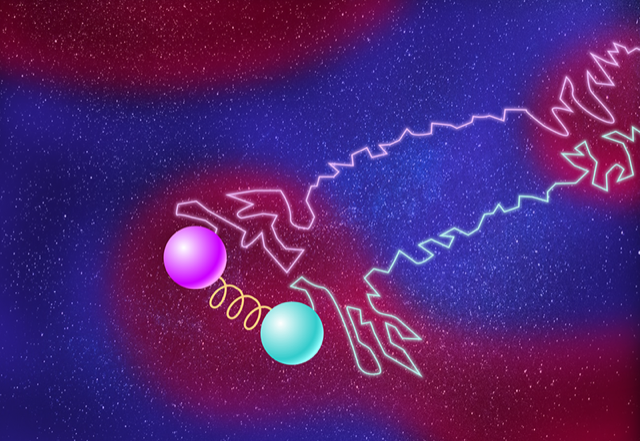
In their study, the duo considered two particles diffusing through a liquid that contains hot and cold patches of various sizes (Fig. 1). Nicoletti and Busiello started by computing the effect of the environment on the particles’ motions. They modeled changes in temperature by randomly switching the temperature around the particles between hot and cool, an action that mimicked motion between patches of hot and cold liquid. They derived the joint probability distribution of the particles’ positions in the asymptotic limit that the temperature patches were large (slow temperature switching). Then they calculated the mutual information between the two particles, finding that it is exactly the Shannon entropy of the fluctuating environment. (In the asymptotic limit of small patches—quick temperature switching—the particles were independent and thus contained no environmental contribution to their mutual information.) Nicoletti and Busiello then repeated the calculations for two spring-coupled particles, finding that the mutual information between them is the superposition of the environmental mutual information for two uncoupled particles and that of the mutual information for interactions intrinsic to two coupled particles. Thus, they were able to separate intrinsic and extrinsic effects.
Being able to bound the effect of the environment on the dynamics of a biological process will aid in teasing out the properties of intrinsic interactions from data, for example in inferring the presence and strength of connections between neurons from voltage recordings. Since Nicoletti and Busiello’s calculations place bounds on the mutual information, rather than giving its definite value, the framework should be applicable to a wide range of models.
Information-theory-based methods have already been applied to solve neuroscience problems, such as reconstructing the connectivity networks of neurons in the brain [4] and mapping the flow of information between regions of the brain [5]. This new work has the potential to extend research in this direction, adding another tool for understanding these complex systems. The approach could also help in developing physically interpretable machine-learning approaches for probing biological systems. In a broad sense, machine-learning techniques take noisy data, which may contain latent variables, and disentangle those data. (Latent variables are like environmental noise in that they are not directly observed.) But machine learning remains a black-box approach, something the new method could help change.
There may also be another impact of Nicoletti and Busiello’s work: understanding the effect of the timescale of the noise. The duo showed that the speed at which the temperature of the environment changed impacted the amount of mutual information between the particles from extrinsic fluctuations. This change in how much the system responded to noise at different timescales points to an emerging picture in biology: Noise is not always a fluctuation about an average that can be ignored or a nuisance that hinders measurement of a parameter. At the proper timescale, noise can be important to the function of a system [6–8]. It is thus important that researchers continue to develop tools to analyze systems that include noise.
References
- W.-X. Wang et al., “Data based identification and prediction of nonlinear and complex dynamical systems,” Phys. Rep. 644, 1 (2016).
- A. Hilfinger and J. Paulsson, “Separating intrinsic from extrinsic fluctuations in dynamic biological systems,” Proc. Natl. Acad. Sci. U.S.A. 108, 12167 (2011).
- G. Nicoletti and D. M. Busiello, “Mutual information disentangles interactions from changing environments,” Phys. Rev. Lett. 127, 228301 (2021).
- D. Zhou et al., “Granger causality network reconstruction of conductance-based integrate-and-fire neuronal systems,” PLOS ONE 9, e87636 (2014).
- B. L. Walker and K. A. Newhall, “Inferring information flow in spike-train data sets using a trial-shuffle method,” PLOS ONE 13, e0206977 (2018).
- B. Walker et al., “Transient crosslinking kinetics optimize gene cluster interactions,” PLOS Comput. Biol. 15, e1007124 (2019).
- B. Walker and K. Newhall, “Numerical computation of effective thermal equilibrium in stochastically switching Langevin systems,” arXiv:2104.13271.
- K. B. Patel et al., “Limited processivity of single motors improves overall transport flux of self-assembled motor-cargo complexes,” Phys. Rev. E 100, 022408 (2019).